In the intricate machinery of the human body, cells continuously exchange massive amounts of information. To communicate, they utilize micro- and nano-sized "courier packages" known as Extracellular Vesicles (EVs, which include exosomes). These vesicles encapsulate proteins and nucleic acids, while their lipid bilayer shells are rich in critical biological information that plays a pivotal role in immune regulation and tumor progression. However, due to their extremely small size (30 to 150 nanometers), capturing them using traditional ultracentrifugation requires substantial sample volumes, consumes excessive time, and often compromises vesicle bioactivity. This has created a longstanding paradox in the industry: balancing the need for "large sample volumes" against the reality of "obtaining rare, trace-level samples."
Recently, a research team led by Professor Shuailong Zhang from the Beijing Institute of Technology, in collaboration with multiple scholars, published an ingenious solution in Analytical Chemistry. They developed a hyphenated Digital Microfluidics-Mass Spectrometry (DMF-MS) platform, establishing a fully automated, miniaturized "exosome sorting and unboxing assembly line." This innovation achieves ultra-fast enrichment and in-situ lipid extraction of EVs from trace, microliter-level samples. Remarkably, the technology condenses the exosome enrichment and lipid extraction processing time from over 2 hours to a mere 15 minutes, enabling in-depth profiling of microscopic samples.
Decoding the Mechanism: Letting Droplets Dance on Chips to Replace Cumbersome Centrifuge Tubes
To shatter the throughput and volume bottlenecks of traditional sample pretreatment, the research team completely discarded test tubes and pipettes, pivoting entirely to Digital Microfluidics (DMF) technology.
The study ingeniously leveraged an Electrowetting-on-Dielectric (EWOD) based DMF platform (whose underlying technology is highly consistent with Optoseeker Biotech's DropletBot® Digital Microfluidic Platform). On these chips, microliter or even nanoliter-sized droplets act as freely maneuverable "chess pieces." By utilizing a programmable underlying electrode array to switch electric fields, the system achieves highly precise control over droplet generation, movement, merging, and splitting (as shown in Figure 1).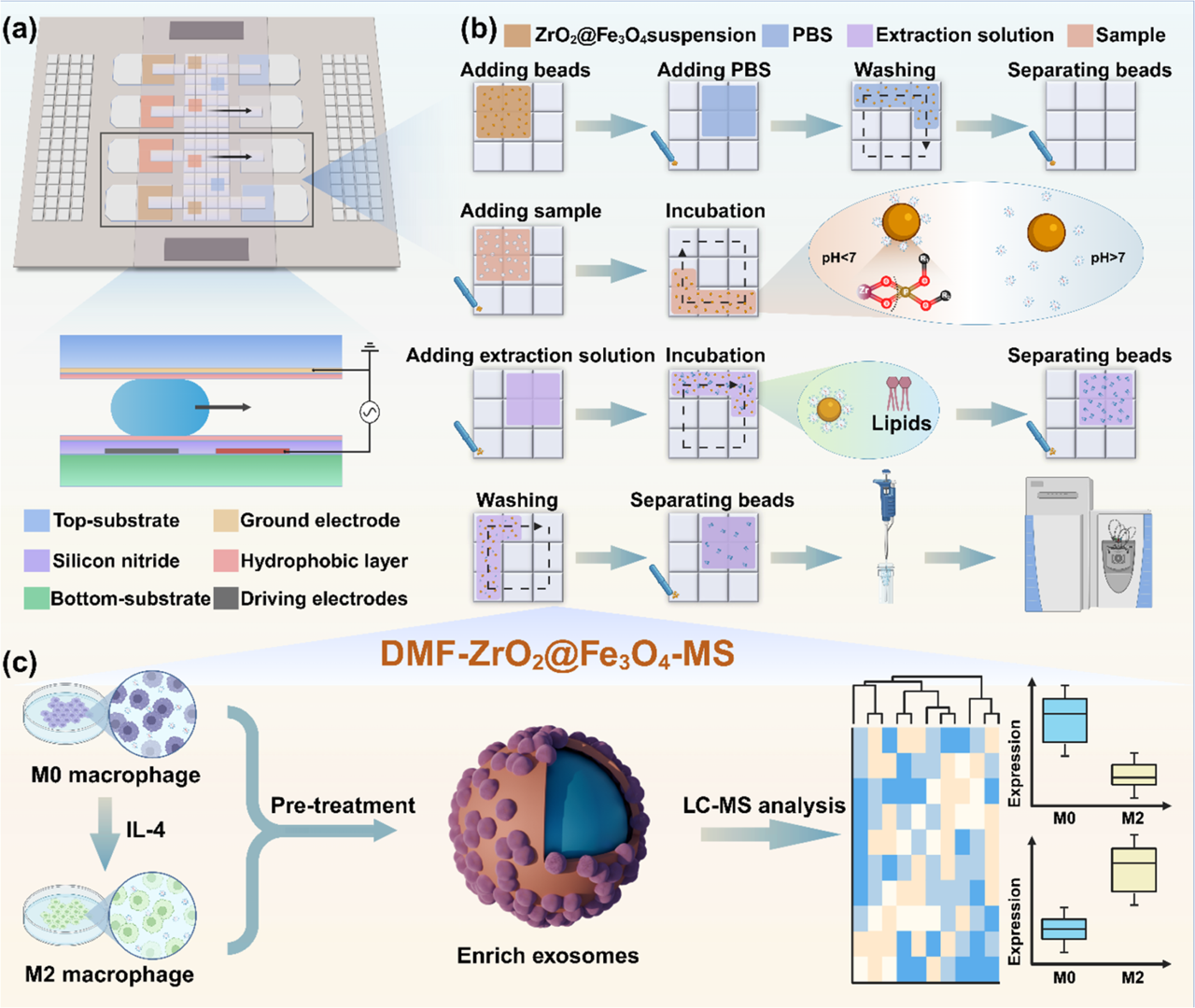
Figure 1. Schematic illustration of EV enrichment and lipid proffling using ZrO2@Fe3O4 on DMF chip. (a) Structure of DMF chip. (b) Detailed workffow for EV enrichment and lipid extraction. (c) All-in-one DMF strategy for sample pretreatment and downstream analysis.
To accurately "fish" for exosomes within these moving droplets, the researchers designed highly specialized "microscopic fishhooks": zirconia-coated magnetic nanoparticles (ZrO2@Fe3O4). The brilliance of this design lies in specific chemical bond recognition. The lipid bilayer surface of exosomes is rich in phosphate groups, while the zirconia surface features Zr4+ Lewis acid sites. When a droplet containing magnetic beads merges with a sample droplet on the chip, the Zr4+ acts like Velcro, rapidly and specifically binding to the phosphate groups on the exosome surface (as shown in Figure 2).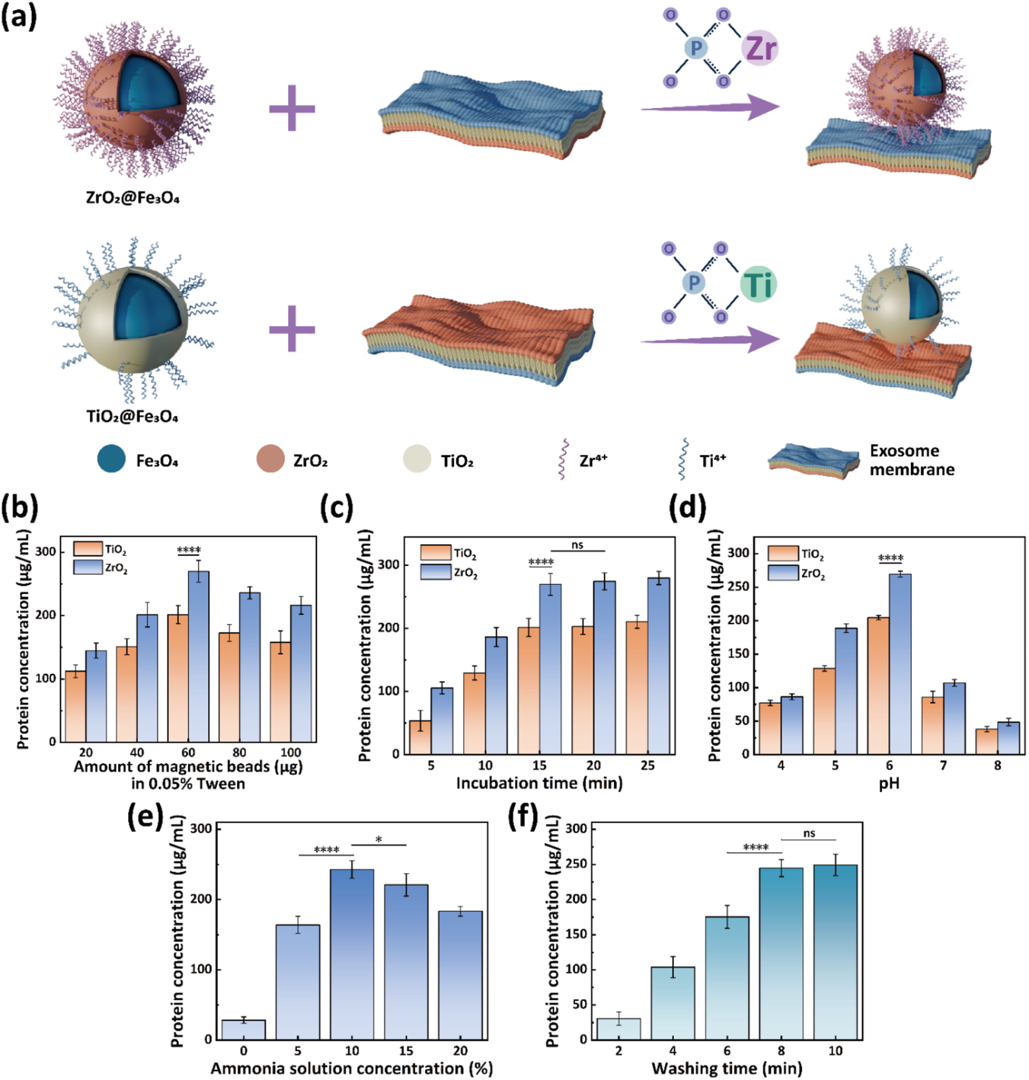
Figure 2. Optimization of EV enrichment and elution conditions for the DMF-ZrO2@Fe3O4 method. (a) Schematic illustration of the chemical mechanism for EV enrichment using ZrO2@Fe3O4 and TiO2@Fe3O4 beads. (b−d) Effects of magnetic beads, incubation time, and solution pH on EV enrichment efffciency. (e, f) Effects of ammonia concentration and elution time on EV elution efffciency. (ns: not signiffcant, *: P < 0.05, ****: P < 0.0001).
Inside this micro-scale "processing plant," the entire closed-loop biochemical reaction—capture, washing, and extraction—is completed entirely within the droplets. Driven by electric fields, droplets containing exosomes and magnetic beads shuttle along pre-set routes to ensure thorough mixing. Subsequently, the system introduces fresh washing droplets to remove impurity proteins, followed by the direct addition of chloroform and acetonitrile mixed droplets on the chip for in-situ lipid extraction. The entire process is enclosed and automated, completely eliminating sample adsorption losses and cross-contamination risks associated with manual pipetting.
Breakthrough Results: 15 Minutes for Deep Lipid Profiling from Micro-Volume (<2.5 μL) Samples
This highly innovative system design yielded striking experimental data. 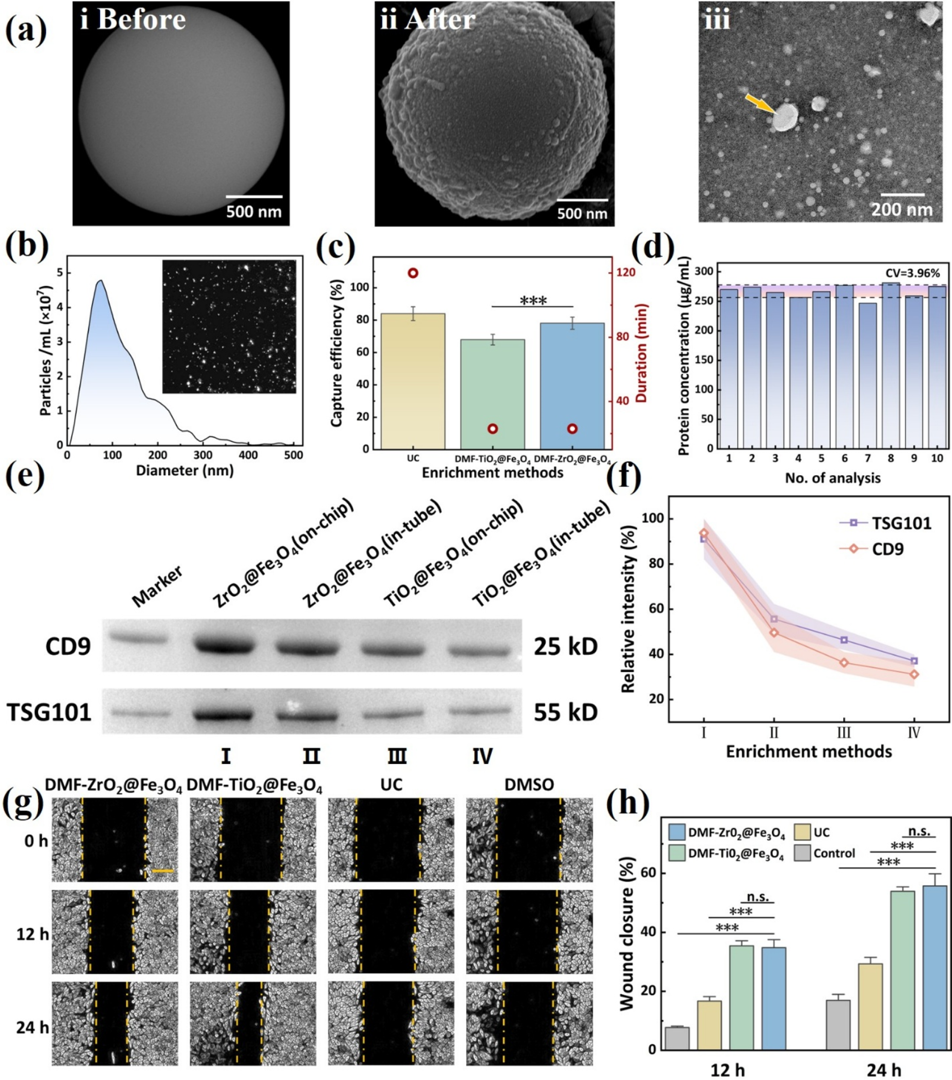
Figure 3. Characterization and evaluation of EVs enriched using the DMF-ZrO2@Fe3O4 strategy. (a) SEM images of ZrO2@Fe3O4 beads before (i) and after (ii) incubation with EV samples on the DMF chip, and a TEM image of the eluted vesicles with an arrow pointing to the cup-shaped EV (iii). (b) NTA of the isolated EVs. (c) Comparison of the recovery rate and time required for EV enrichment using different approaches. (d) Reproducibility analysis of EV enrichment using the DMF-ZrO2@Fe3O4 method. (e, f) Western blot analysis of CD9 and TSG10l in EVs isolated either on-chip or intube using ZrO2@Fe3O4 or TiO2@Fe3O4. (g) EVs enriched with different methods were added to the medium for culturing HeLa cells in a scratch assay. The control group was treated with an EV-free culture medium. Cell migration was monitored at 12 and 24 h of culture. (The scale bar is 200 μm.) (h) The scratch healing rate of HeLa cells was determined by calculating the ratio of the scratch width at 12 or 24 h to the scratch width at 0 h. (n.s.: not signiffcant, ***: P < 0.001).
By directly transferring the extracted droplets from the chip to Mass Spectrometry (MS) analysis, the research team successfully identified 349 lipid molecules in HeLa cell-derived exosomes, nearly a hundred more than the traditional Bligh-Dyer extraction method. These lipids encompass multiple categories, including glycerophospholipids, sphingolipids, and glycerolipids, robustly proving the immense coverage capability of this miniaturized platform for non-targeted lipidomics analysis (as shown in Figure 4).
Figure 4. Lipid proffling of HeLa-cell-derived EVs. (a, b) Massspectra of extracted exosomal lipidsin positive and negative ion modes. (c) Comparison of lipid identiffcation quantities between on-chip extraction and the classic Bligh−Dyer method. (d) Classiffcation and (e) subclassiffcation of identiffed lipids from enriched EVs from HeLa cells. (f) Correlation analysis of lipid abundances between replicates.
Deciphering the "Peacekeeping" Code of Macrophages
To validate the technology's performance in complex real-world scenarios, researchers applied it to a highly challenging immunological subject: exosome lipid profiling of macrophage polarization states.
Macrophages are the "scavengers" of the immune system, primarily divided into the resting M0 phenotype and the M2 phenotype, which boasts anti-inflammatory and tissue repair functions. However, isolating specific polarization states and their secretions is extremely difficult, making sample volumes incredibly scarce.
Using this novel DMF-MS platform, the team conducted precise lipid profile comparisons between M0 and M2 macrophage exosomes. The data unveiled a fascinating biological phenomenon: compared to resting M0 macrophages, M2 macrophage-secreted exosomes exhibited a highly significant upregulation of anti-inflammatory monounsaturated fatty acids (like palmitoleic acid) and Omega-3 fatty acids (like DHA); meanwhile, cholesteryl esters—the intracellular lipid storage form—decreased drastically (as shown in Figure 5).
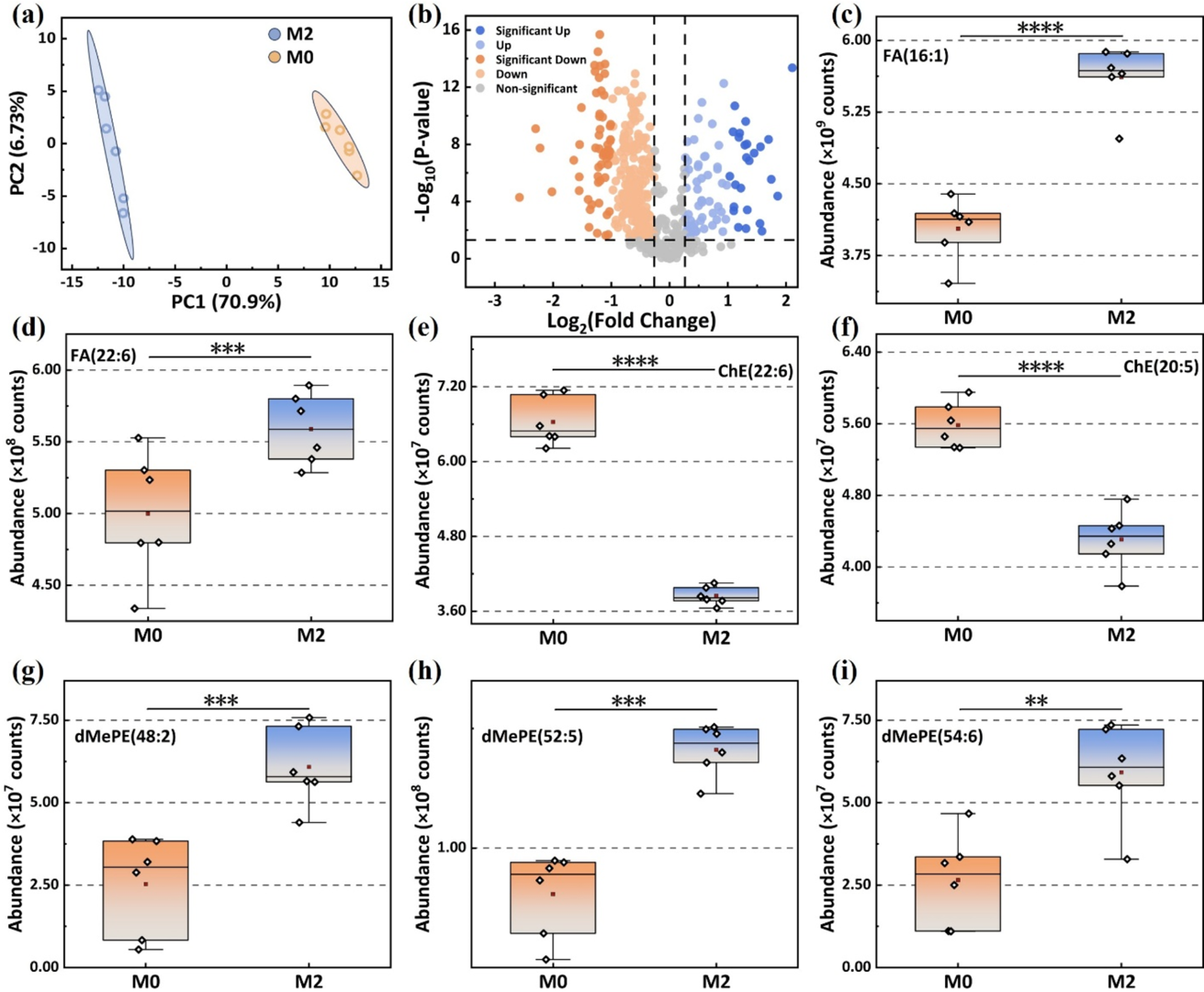
Figure 5. Lipid analysis of EVs derived from M0 and M2 mouse macrophages. (a, b) Principal component analysis (PCA) and volcano plot of lipids in EVs from M0 and M2 mouse macrophages (Up: FC > 1.5, P < 0.05, Signiffcant Up: FC > 2, P < 0.05). Abundance comparisons of FA(16:1) (c), FA(22:6) (d), ChE(22:6) (e), ChE(20:5) (f), dMePE(48:2) (g), dMePE(52:5) (h), and dMePE(54:6) (i) in EVs from M0 and M2 mouse macrophages. (**: P < 0.01,***: P < 0.001,****: P < 0.0001.).
This micro-level lipid remodeling perfectly mirrors the physiological "inflammation-calming and repair-promoting" macroscopic role of M2 macrophages. This not only provides fresh molecular evidence for understanding immuno-metabolic mechanisms but directly proves the platform's irreplaceable superiority in handling rare cell samples.
A Microscopic Perspective for Future Analytical Paradigms
This work opens a new window for deep molecular characterization of trace clinical samples.
For researchers and clinical diagnostics professionals, condensing the complex exosome isolation and lipid extraction processes onto a card-sized microfluidic chip is far beyond a mere engineering flex. It directly solves the core pain points of traditional workflows: large dead volumes, high sample loss, and human operation variability. With the popularization of digital microfluidic hardware and the optimization of underlying algorithms, this highly efficient, high-fidelity "drop of blood" miniaturized workflow is poised to become a mainstream, standard tool in immuno-metabolism research, rare disease biomarker screening, and liquid biopsy.
Featured Product Integration: 
Optoseeker Biotech's DropletBot® Digital Microfluidic Platform is an automated system powered by Electrowetting-on-Dielectric (EWOD) based Digital Microfluidics (DMF). By altering the contact angle of droplets on a hydrophobic surface, it precisely maneuvers discrete droplets across an electrode array. It is broadly applicable in biological and chemical fields, including diagnostics, cell assays, and chemical synthesis. Featuring a modular design and an intuitive user interface, DropletBot® can drive multiple independent channels and provides dynamic impedance sensing for real-time measurements of droplet position, velocity, and electrostatic forces.
Click to learn more about the product
Original Publication:
Streamlined Digital Microffuidics-Mass Spectrometry Strategy for Extracellular Vesicle Enrichment and Lipid Proffling
DOI.org/10.1021/acs.analchem.5c07527







